

This story is adapted from our recent feature, Of Mosquitoes and Models: Tracking Disease by Satellite.
In 2017, Yemen experienced one of its worst cholera outbreaks on record. Following heavy rains, flooding, and mass movement of the population due to civil unrest, more than one million people were suspected of contracting cholera and at least 2,000 died. A few scientists saw it coming, and they are now working to make sure people are prepared for future cholera outbreaks in Yemen and around the world.
Cholera is a waterborne bacterial infection that can spread quickly through a population. The disease is primarily contracted by consuming water or food contaminated with the cholera bacteria, Vibrio cholerae. It causes uncontrollable diarrhea that, if left untreated, can result in dehydration or death.
A team of NASA-funded researchers has been using satellite and ground-based data to forecast the risk of cholera in Yemen and other countries. The map above shows the forecasted risk of cholera in Yemen from August 10 to September 6, 2020. It was created with the Cholera Prediction Modeling System, which incorporates NASA precipitation data, air temperature data from NASA’s MERRA-2 reanalysis product, and population data. The number of cholera cases could increase in coming weeks, influenced by heavy rains that usually fall in August, though researchers predict the outbreaks should be limited to a few hotspots unless there is a large population displacement.
Led by hydrologist Antar Jutla of the University of Florida, the Cholera Prediction Modeling System got its first real-world test in 2017. Observing heavy rain and warm temperatures, Jutla and colleagues predicted that a cholera outbreak could occur in Yemen that summer. However, the team had concerns about the accuracy of their model because they could not get reliable health data from the war-torn country. They made their prediction privately, but were not able to share it with Yemeni officials.
Yemen proceeded to have a devastating cholera outbreak that June. Jutla’s model achieved 92 percent accuracy in predicting areas where cholera appeared that year. It even identified outbreaks in inland areas that are not usually susceptible to the disease. Following that successful prediction, Jutla and colleagues partnered with UNICEF and the UK Department for International Development to share weekly reports on potential outbreaks in Yemen in order to guide the delivery of prevention and relief resources.
“This 2017 outbreak was a rare occurrence in Yemen, but the model was able to forecast it,” said John Haynes, a program manager in NASA’s Earth Applied Sciences Program, which funds Jutla. “It was a really big breakthrough to learn that the model could be moved to different parts of the world and still work well.”
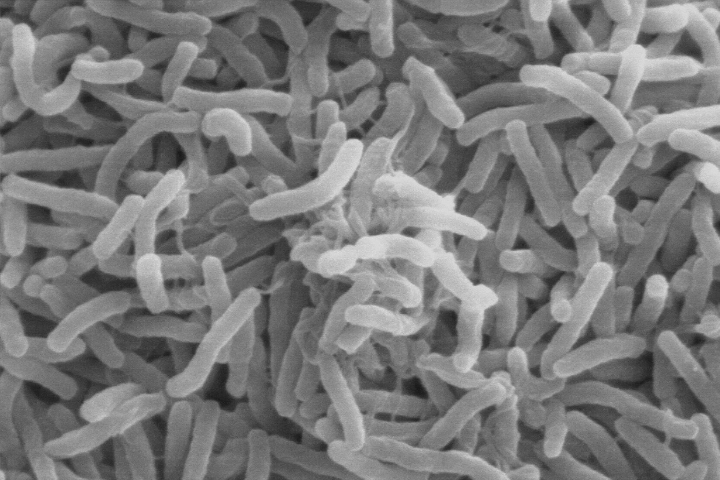
The idea that cholera outbreaks can be tracked partly by looking at specific environmental conditions is relatively new. In the 1960s, University of Maryland biologist Rita Colwell made the discovery that Vibrio cholerae bacteria can live in the guts of microscopic aquatic animals called copepods. Further investigations showed that disease-carrying copepods could lurk in the coastal ocean for months to years. Other research showed that when the right environmental conditions arise, Vibrio cholerae can show up in water supplies and spread disease to people. But at the time researchers did not know which environmental factors brought the bacteria-carrying copepods into water supplies.
In 2011, Jutla began working with Colwell to understand why cholera outbreaks occur. After analyzing decades of cholera outbreak and satellite data in various countries, the team found that cholera outbreaks took two different forms: endemic and epidemic. Endemic disease means there is a cyclical or seasonal recurrence because the bacteria is constantly present in the environment, even if at a low level.
The second form—the epidemic mode—occurs suddenly and sporadically. These outbreaks typically occur after a disaster (such as flooding) contaminates clean water sources or damages water infrastructure. They also alter normal human behavior; for instance, causing populations to relocate suddenly. These damages and disruptions often increase the chance that the infectious bacteria will enter and contaminate local water sources.
Epidemic episodes tend to result in more deaths than endemic episodes because the population is often not prepared for the sudden outbreak and subsequently leads to massive infection rates. However, sometimes sustained epidemic incidents can keep the Vibrio cholerae bacteria in the environment for a long time and cholera becomes a regular occurrence—which is what happened in Yemen.
Yemen never fully recovered from its 2017 cholera outbreak, which introduced Vibrio cholerae bacteria back into the region. Since then, poor sanitation systems have allowed the bacteria to persist in drinking water. There are more occurrences of cholera now, particularly in coastal regions where salty rivers create conditions for the bacteria to thrive.
According to UNICEF, Yemen had more than 2 million suspected cases between April 2017 to June 2020. What seemed like a one-time epidemic is now turning endemic, and Jutla and Colwell are working to understand conditions that allow the infectious bacteria to stick around in a human population.
For Yemen, the team noted that if water sanitation systems are not improved, the number of cases may become so high that cholera may occur seasonally for years. Colwell and Jutla have recently been working with the UK’s Meteorological Office and its Department for International Development to develop protocols to better educate people on how to interpret the model’s results and to reassess prevention and education methods.
“Ultimately, cholera can only be controlled if all the relevant humanitarian agencies work in tandem and synchrony so that we can make this disease a thing of past,” said Jutla.
NASA Earth Observatory map by Lauren Dauphin using data from Antar Jutla and Moiz Usmani, University of Florida. Scanning electron microscope image courtesy of Kirn et al., Dartmouth Medical School, via Wikimedia Commons. Story by Kasha Patel.